Examples
#include "numerics.hpp"
int main() {
const double sigma = 10.0, rho = 28.0, beta = 8.0 / 3.0;
auto lorenz = [&](double, const num::Vector& s, num::Vector& ds) {
ds[0] = sigma * (s[1] - s[0]);
ds[1] = s[0] * (rho - s[2]) - s[1];
ds[2] = s[0] * s[1] - beta * s[2];
};
// simulation observer function to store (x, z) each step
num::Series xz;
auto observer = [&](double, const num::Vector& y) {
xz.store(y[0], y[2]);
};
num::solve(
num::ODEProblem{lorenz, {1.0, 0.0, 0.0}, 0.0, 50.0},
num::RK45{.rtol = 1e-8, .atol = 1e-10, .max_steps = 2000000},
observer);
num::plt::plot(xz);
num::plt::title("Lorenz attractor (sigma=10, rho=28, beta=8/3)");
num::plt::xlabel("x");
num::plt::ylabel("z");
num::plt::savefig("lorenz.png");
}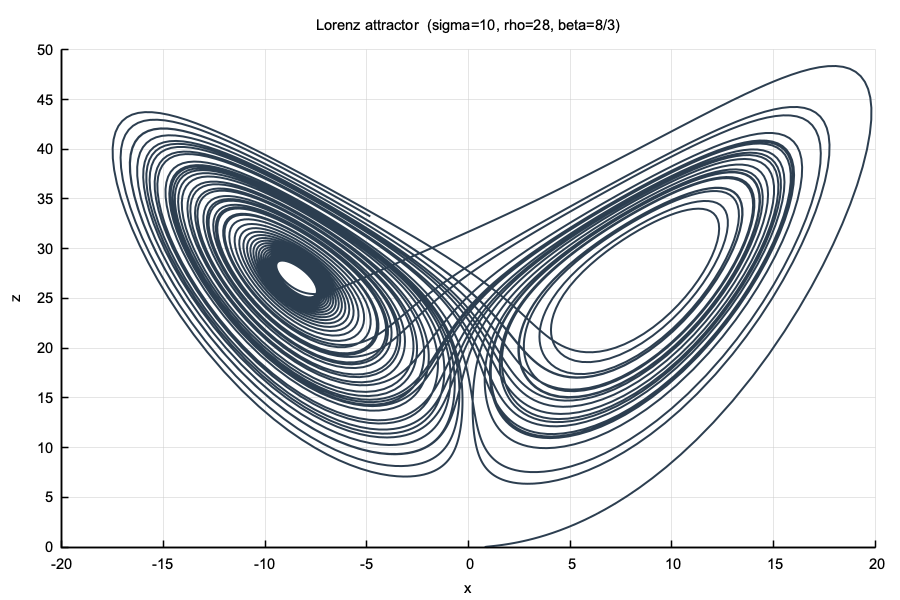
num::solve + num::RK45#include "plot/plot.hpp"
#include "spectral/fft.hpp"
int main() {
const int N = 1024; const double fs = 1000.0; // N samples at fs Hz
num::Vector x(N); // real-valued time-domain signal
for (int i = 0; i < N; ++i)
x[i] = std::sin(2*M_PI*50.0*i/fs)
+ 0.5*std::sin(2*M_PI*120.0*i/fs);
// rfft output has N/2+1 bins: real input has conjugate symmetry,
// so only the non-redundant half [0 .. N/2] is stored
num::CVector X(N/2 + 1); // complex frequency-domain coefficients
num::spectral::rfft(x, X);
num::Series signal, spectrum; // lists of (x, y) pairs for plotting
for (int i = 0; i < 100; ++i)
signal.emplace_back(i/fs*1000.0, x[i]); // first 100 ms
for (int k = 0; k < (int)X.size(); ++k)
spectrum.emplace_back(k*fs/N, std::abs(X[k])); // bin k maps to freq k*fs/N Hz
num::plt::subplot(2, 1);
num::plt::plot(signal);
num::plt::title("Signal: 50 Hz + 0.5 * 120 Hz");
num::plt::xlabel("time (ms)"); num::plt::ylabel("amplitude");
num::plt::next();
num::plt::plot(spectrum);
num::plt::title("FFT power spectrum");
num::plt::xlabel("frequency (Hz)"); num::plt::ylabel("|X[k]|");
num::plt::savefig("fft_demo.png");
}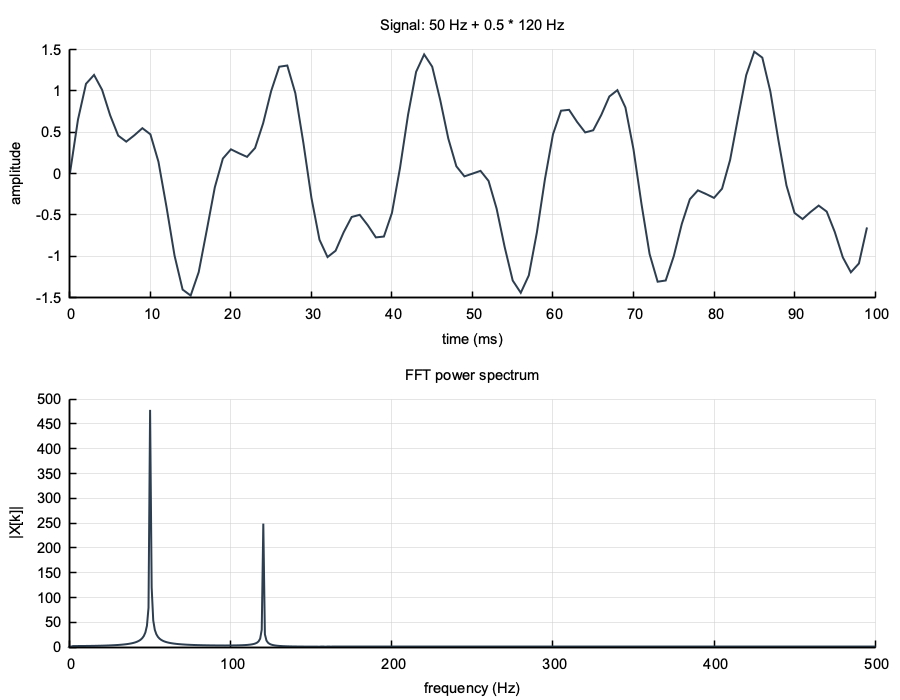
num::spectral::rfft; peaks at 50 Hz and 120 Hz#include "numerics.hpp"
#include "spatial/pbc_lattice.hpp"
#include "stochastic/boltzmann_table.hpp"
int main() {
const int N = 64, NN = N*N;
num::PBCLattice2D nbr(N);
std::vector<double> spins(NN);
std::mt19937 rng(42);
num::Series curve;
for (double T = 0.5; T <= 4.01; T += 0.1) {
std::fill(spins.begin(), spins.end(), 1.0);
double beta = 1.0 / T;
auto accept = [&](int i) {
double ns = spins[nbr.up[i]] + spins[nbr.dn[i]]
+ spins[nbr.lt[i]] + spins[nbr.rt[i]];
return num::markov::boltzmann_accept(2.0*spins[i]*ns, beta);
};
auto flip = [&](int i) { spins[i] = -spins[i]; };
auto measure = [&]() {
return std::abs(std::accumulate(spins.begin(), spins.end(), 0.0) / NN);
};
double m = num::solve(
num::MCMCProblem{accept, flip, NN},
num::Metropolis{.equilibration=2000, .measurements=500},
measure, rng);
curve.store(T, m);
}
num::plt::plot(curve);
num::plt::xlabel("T"); num::plt::ylabel("|<m>|");
num::plt::savefig("ising_demo.png");
}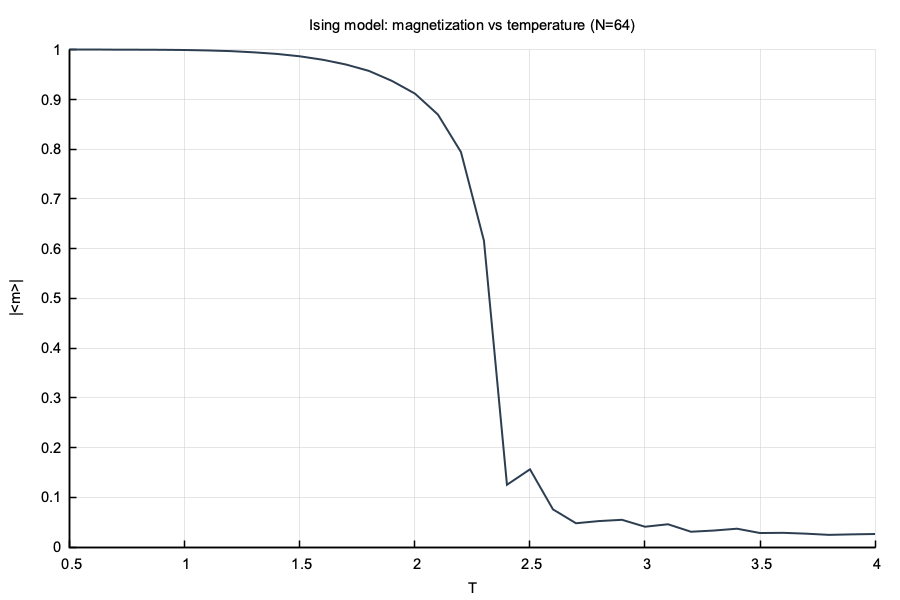
num::solve + num::Metropolis; |m| drops at Tc ≈ 2.27#include "numerics.hpp"
#include <string>
using namespace num;
int main() {
const int N = 64;
const double kappa = 1.0, sigma = 0.06, T_end = 0.012;
const double h = 1.0 / (N + 1);
const double dt = 4.0 * h * h; // 16x larger than explicit limit
const int nstep = static_cast<int>(T_end / dt) + 1;
const double coeff = kappa * dt / (h * h);
// A = I - coeff*L (sparse SPD -- CG converges in O(N) iterations)
Grid2D grid{N, h};
SparseMatrix A = pde::backward_euler_matrix(grid, coeff);
LinearSolver solver = make_cg_solver(A);
ScalarField2D u0(grid, [=](double x, double y) {
return gaussian2d(x, y, 0.5, 0.5, sigma);
});
ScalarField2D u = u0;
solve(u, BackwardEuler{.solver=solver, .dt=dt, .nstep=nstep});
plt::subplot(1, 2);
plt::heatmap(u0); plt::title("t = 0");
plt::next();
plt::heatmap(u); plt::title("t = " + std::to_string(T_end).substr(0, 6));
plt::savefig("heat_demo.png");
}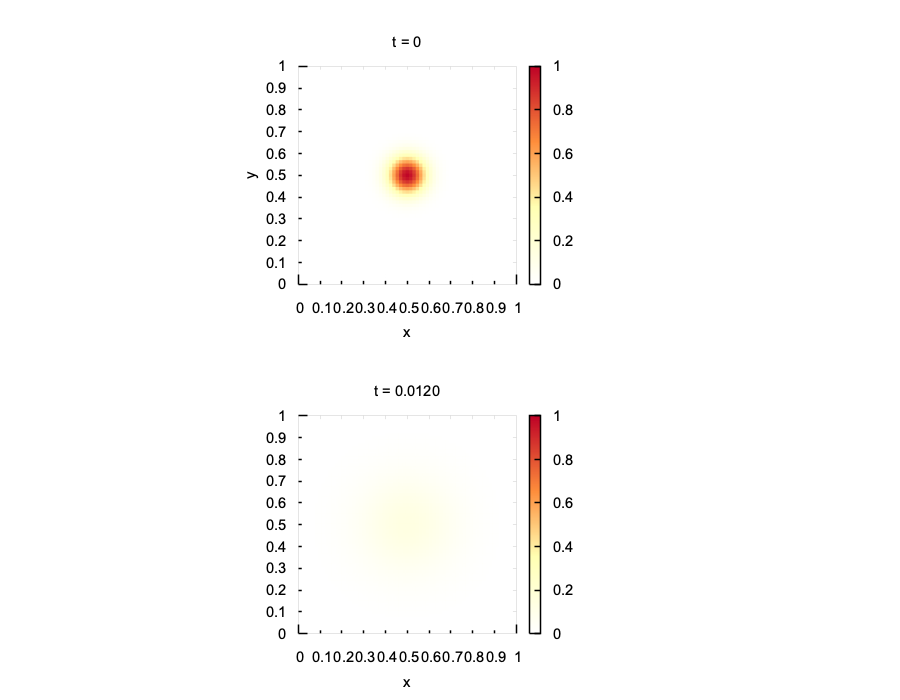
num::solve + num::BackwardEuler