Stochastic Simulation of Biochemical Networks
Finite State Projection methods for the Chemical Master Equation
The Chemical Master Equation (CME) governs the time-evolution of the probability distribution over all possible molecule counts in a stochastic reaction network:
\[\frac{dP(\mathbf{x},t)}{dt} = \sum_{\mathbf{x}'\neq\mathbf{x}} \Bigl[W_{\mathbf{x}'\to\mathbf{x}}\,P(\mathbf{x}',t) - W_{\mathbf{x}\to\mathbf{x}'}\,P(\mathbf{x},t)\Bigr]\]The state space \(\mathbf{x} \in \mathbb{N}^N\) is in principle infinite, making direct numerical solution intractable. The Finite State Projection (FSP) method truncates the generator matrix:
\[\mathbf{W} = \begin{pmatrix} \mathbf{Q} & \mathbf{R} \\ \mathbf{T} & \mathbf{U} \end{pmatrix} \quad \approx \quad \widetilde{\mathbf{W}} = \begin{pmatrix} \mathbf{Q} & \mathbf{R} \\ \mathbf{0} & \mathbf{0} \end{pmatrix}\]My research focuses on dynamically controlling the truncated state space over time, so the probability mass stays well-resolved without over-allocating memory. The example below shows an FSP simulation of a genetic toggle switch — two mutually repressing genes that can exist in two stable states:
\[\varnothing \xrightarrow{\frac{\beta_A}{1+(B/K_B)^{n_B}}} A,\; A \xrightarrow{\delta_A} \varnothing \qquad \varnothing \xrightarrow{\frac{\beta_B}{1+(A/K_A)^{n_A}}} B,\; B \xrightarrow{\delta_B} \varnothing\]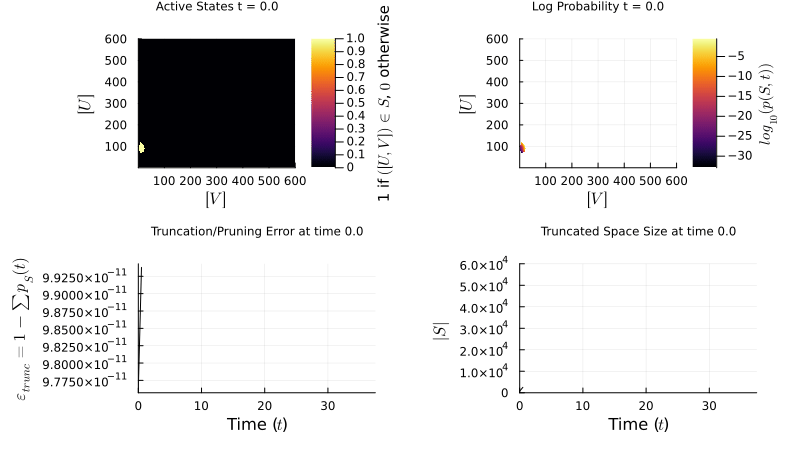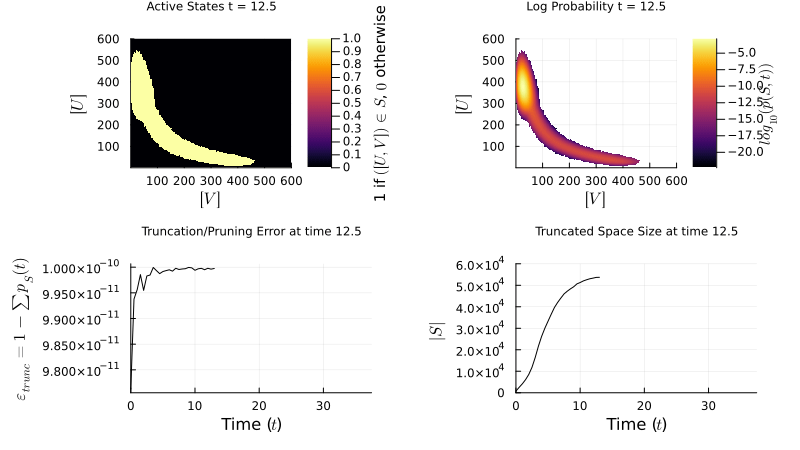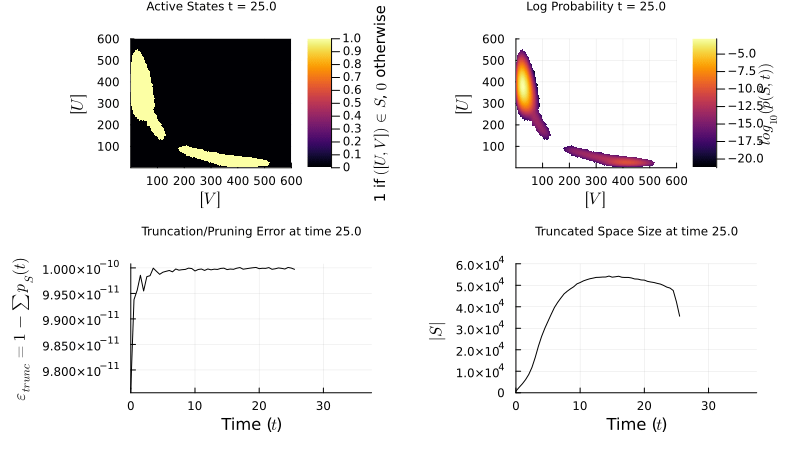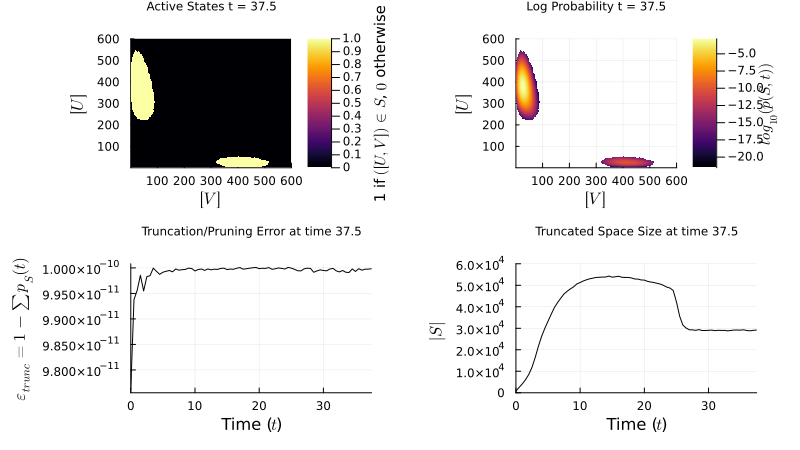
FSP simulation of the genetic toggle switch. The probability distribution evolves over the 2D state space (copy counts of protein A vs. B), revealing bistability.