Koopman Operator Analysis of Reaction Networks
EDMD-based rate recovery from stochastic trajectory data
Rather than evolving probability distributions forward (the CME direction), the Koopman generator acts on observables \(f: \mathbb{N}^N \to \mathbb{R}\):
\[\frac{d}{dt}\langle f \rangle = \left\langle \sum_{k=1}^M \alpha_k(\mathbf{x})\bigl[f(\mathbf{x}+\boldsymbol{\zeta}_k) - f(\mathbf{x})\bigr]\right\rangle = \langle \mathcal{A} f \rangle\]where \(\alpha_k(\mathbf{x}) = c_k \cdot \psi_{m(k)}(\mathbf{x})\) are the reaction propensities. Since each propensity factors as a known polynomial \(\psi_{m(k)}\) times an unknown rate constant \(c_k\), the Koopman generator is linear in the rates:
\[\mathcal{A} = \sum_{k=1}^M c_k \mathcal{A}_k\]Using Extended Dynamic Mode Decomposition (EDMD), I recover the dominant Koopman eigenfunctions and eigenvalues directly from SSA trajectory data — without ever forming the full CME generator. The dominant modes capture the slow timescales of the network, and the rate constants follow from a linear regression in the Koopman basis.
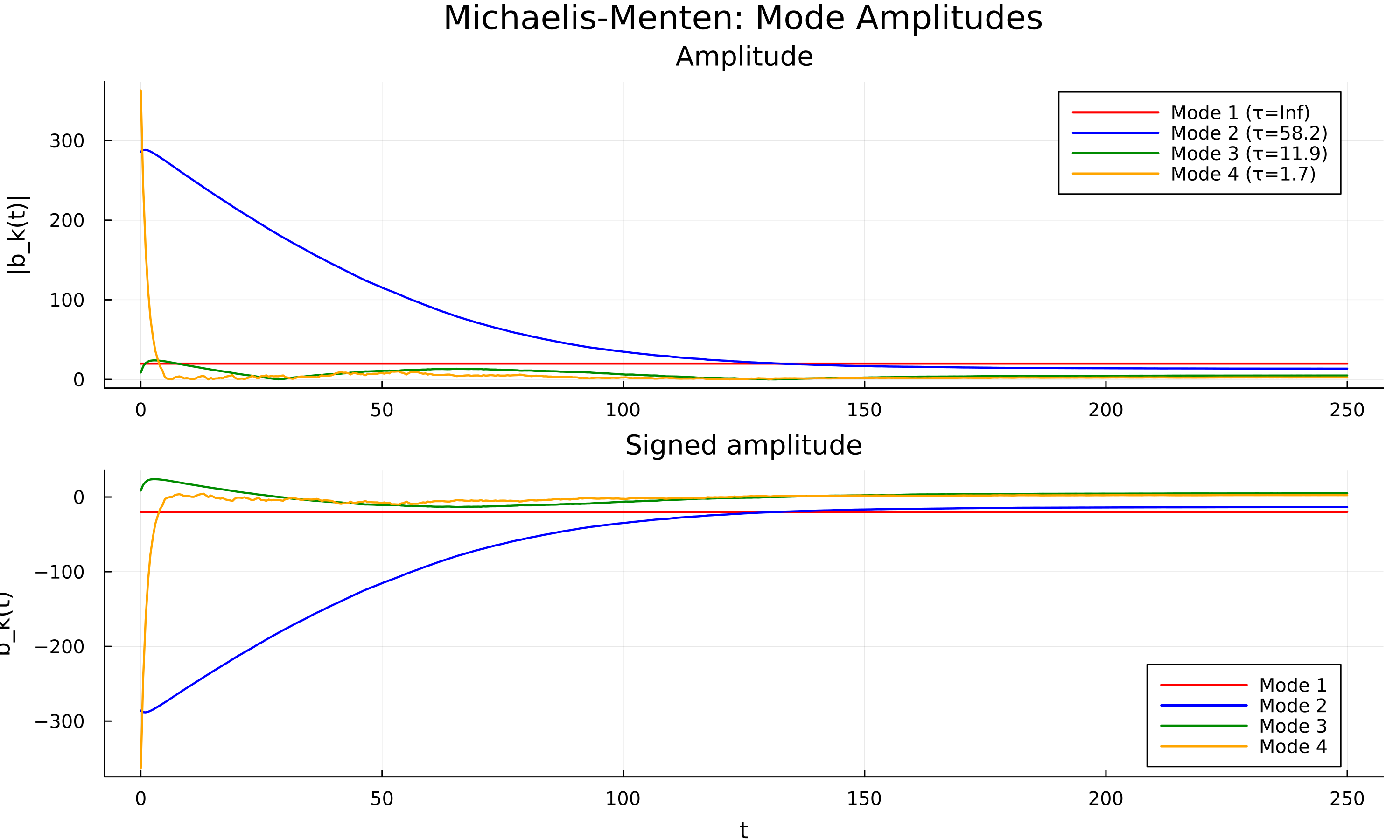
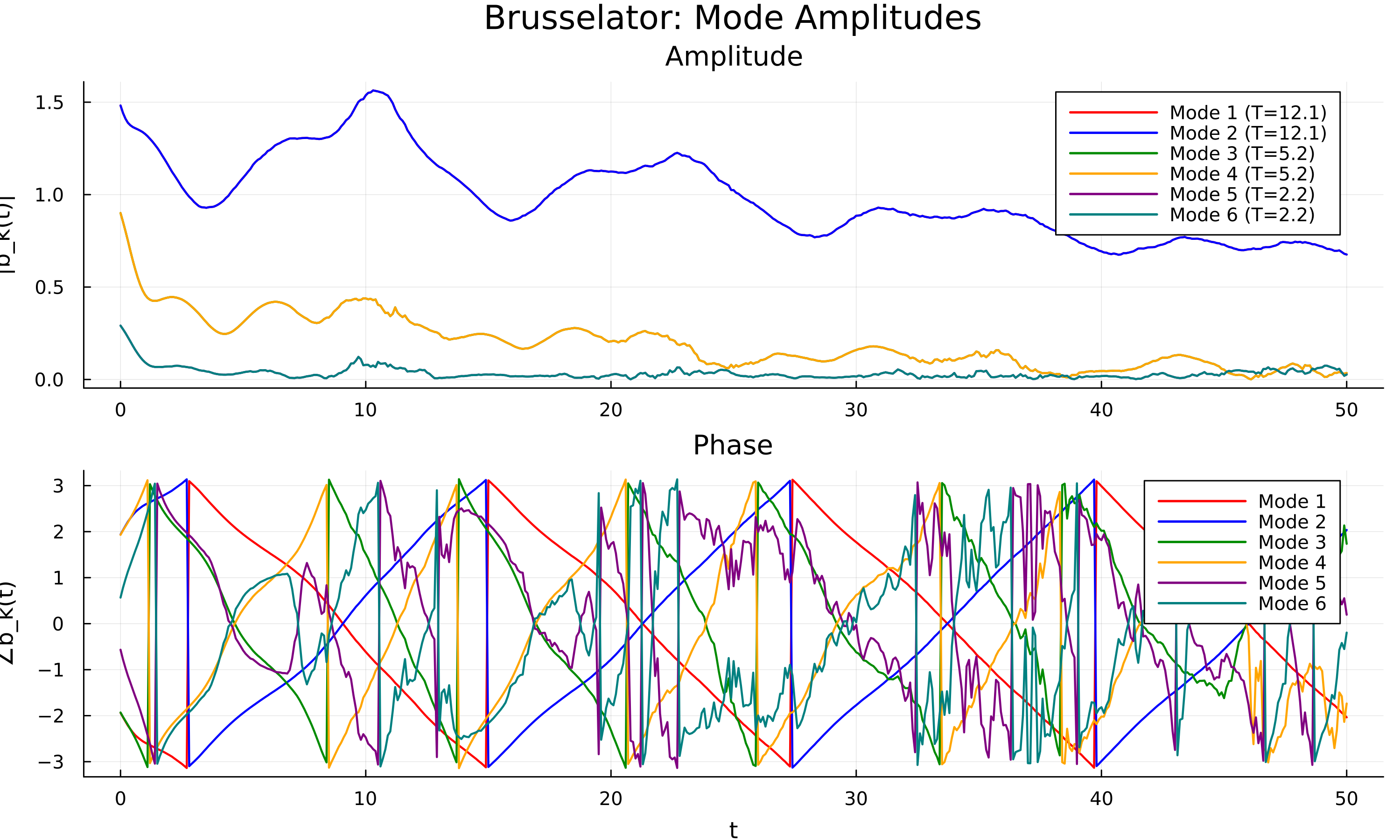
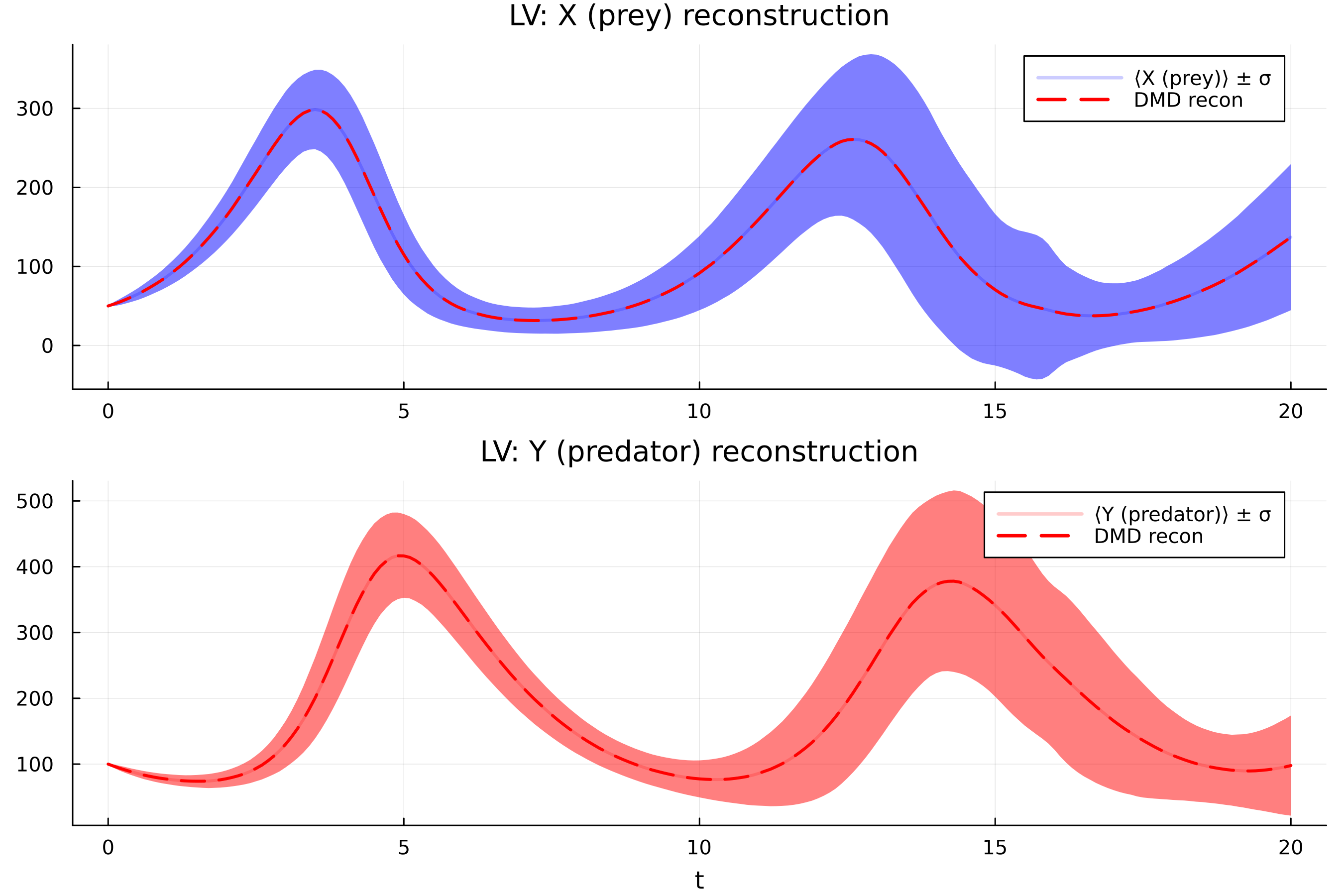
Results on three benchmark networks are shown below.